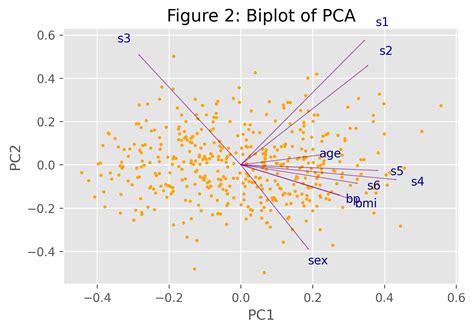
In the realm of data science and machine learning, Principal Component Analysis (PCA) stands as a crucial tool for dimensionality reduction and feature extraction. By transforming complex, high-dimensional data into a more manageable form, PCA allows for better visualization and easier computation. In this comprehensive guide, we will delve into how to master PCA using the scikit-learn library, highlighting practical insights and real-world applications.
Key Insights
- PCA enables simplification of high-dimensional datasets, making them easier to visualize and work with.
- The scikit-learn library offers an efficient implementation of PCA with flexible parameters for customization.
- Implementing PCA through scikit-learn enhances model performance and reduces overfitting.
Understanding the Mechanics of PCA
Principal Component Analysis operates on the principle of finding new orthogonal axes (principal components) that maximize variance in a dataset. This is achieved through an eigen-decomposition of the covariance matrix or singular value decomposition (SVD) of the dataset. By projecting the original data onto these new axes, PCA identifies the directions where the data variation is most significant, allowing for a reduced number of features while retaining most of the information. This technique is especially useful in cases where multicollinearity is a concern or when computational efficiency is paramount.Implementation of PCA with scikit-learn
Scikit-learn’s PCA implementation is both straightforward and highly effective. To begin with, one needs to import the PCA module from scikit-learn and initialize it with desired parameters. For example, then_components parameter defines how many principal components should be retained.
Here's an example:
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
principalComponents = pca.fit_transform(X)In this snippet, `X` represents your dataset. The call to `fit_transform` fits the PCA model to the data and immediately transforms it, reducing it to two principal components.
Advanced Customizations with PCA
The scikit-learn PCA class provides several customizable parameters that can be tailored to fit specific use cases. Thewhiten parameter, for instance, allows you to standardize the components to unit variance. This is particularly useful when the variance in different features differs vastly.
Another useful parameter is svd_solver, which determines the method used to compute the SVD. Depending on the dataset size and shape, different solvers like ‘auto’, ‘full’, ‘randomized’, and ‘arpack’ might offer different performance characteristics. For smaller, dense datasets, the ‘full’ solver may be optimal. In contrast, larger, sparse datasets might benefit from the ‘randomized’ solver due to its computational efficiency.
What is the impact of PCA on model performance?
PCA can significantly improve model performance by reducing overfitting. By limiting the number of features, it mitigates the risk of including noise and multicollinearity, which can degrade model performance. This is especially beneficial in high-dimensional datasets where many features provide little to no additional predictive power.
Can PCA be applied to any type of data?
PCA is most effective on linearly correlated datasets. While it can be applied to any dataset, its benefits are most pronounced in situations where linear relationships dominate. For highly non-linear datasets, alternative dimensionality reduction techniques like t-distributed Stochastic Neighbor Embedding (t-SNE) might yield better results.
PCA, when leveraged correctly, can vastly simplify complex datasets while preserving the intrinsic structure within the data. With scikit-learn, this process becomes not only accessible but also efficient and easy to implement. Whether for visualization, preprocessing, or feature extraction, PCA stands out as an indispensable tool in the data scientist’s toolkit.